Problems tagged with "lecture-06"
Problem #058
Tags: pca, projection, quiz-04, dimensionality reduction, lecture-06
Suppose the direction of maximum variance in a centered data set is
Let \(\vec x = (2, 4)^T\) be a centered data point.
Reduce \(\vec x\) to one dimension by projecting onto the direction of maximum variance. What is the new feature \(z\) obtained from this projection?
Solution
\(z = 3\sqrt 2\).
The projection onto the direction of maximum variance is given by the dot product with \(\vec u\):
Problem #059
Tags: pca, projection, quiz-04, dimensionality reduction, lecture-06
Suppose the direction of maximum variance in a centered data set is
Let \(\vec x = (2, 1, 2)^T\) be a centered data point.
Reduce \(\vec x\) to one dimension by projecting onto the direction of maximum variance. What is the new feature \(z\) obtained from this projection?
Solution
\(z = 3\).
The projection onto the direction of maximum variance is given by the dot product with \(\vec u\):
Problem #060
Tags: pca, projection, quiz-04, dimensionality reduction, lecture-06
Suppose the direction of maximum variance in a centered data set is
Let \(\vec x = (3, 1, -1, 5)^T\) be a centered data point.
Reduce \(\vec x\) to one dimension by projecting onto the direction of maximum variance. What is the new feature \(z\) obtained from this projection?
Solution
\(z = 4\).
The projection onto the direction of maximum variance is given by the dot product with \(\vec u\):
Problem #061
Tags: pca, centering, covariance, quiz-04, lecture-06
Center the data set:
Solution
First, compute the mean:
Then subtract the mean from each data point:
Problem #062
Tags: pca, centering, covariance, quiz-04, lecture-06
Center the data set:
Solution
First, compute the mean:
Then subtract the mean from each data point:
Problem #063
Tags: quiz-04, covariance, lecture-06
Consider the dataset of four points in \(\mathbb R^2\) shown below:
Calculate the sample covariance matrix.
Solution
First, compute the mean:
Then form the centered data matrix \(Z\), whose rows are the centered data points:
The sample covariance matrix is:
Problem #064
Tags: quiz-04, covariance, lecture-06
Consider the data set in the image shown below:
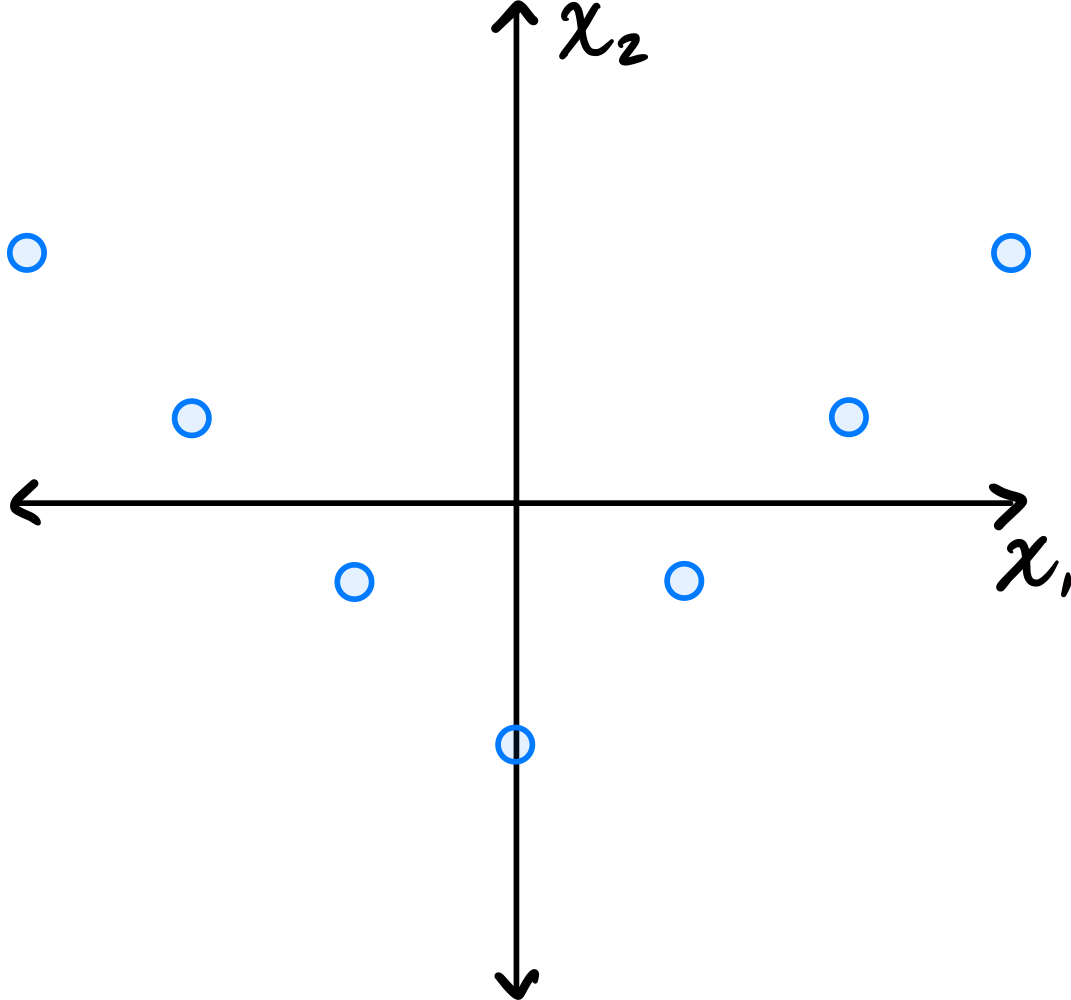
You may assume that this data is already centered and that the symmetry over the \(x_2\) axis is exact.
Which one of the following is true about the \((1, 2)\) entry of the data's sample covariance matrix?
Consider the \((1, 2)\) entry of the covariance matrix, which represents the covariance between the first and second features.
Is the \((1, 2)\) entry of the covariance matrix positive, negative, or zero?
Solution
It is zero.
Because of the symmetry over the \(x_2\) axis, for every point \((a, b)\) in the data set, there is a corresponding point \((-a, b)\). When computing the \((1, 2)\) entry of the covariance matrix, we sum \(\tilde{x}_1^{(i)}\cdot\tilde{x}_2^{(i)}\) over all points. For the symmetric pairs \((a, b)\) and \((-a, b)\), these contributions are \(ab\) and \(-ab\), which cancel out. Therefore, the \((1, 2)\) entry is zero.
Problem #065
Tags: quiz-04, covariance, lecture-06
Let \(\vec{x}^{(1)}, \ldots, \vec{x}^{(6)}\) be a data set of \(6\) points in \(\mathbb R^3\). Shown below are scatter plots of each pair of coordinates (pay close attention to the axis labels):
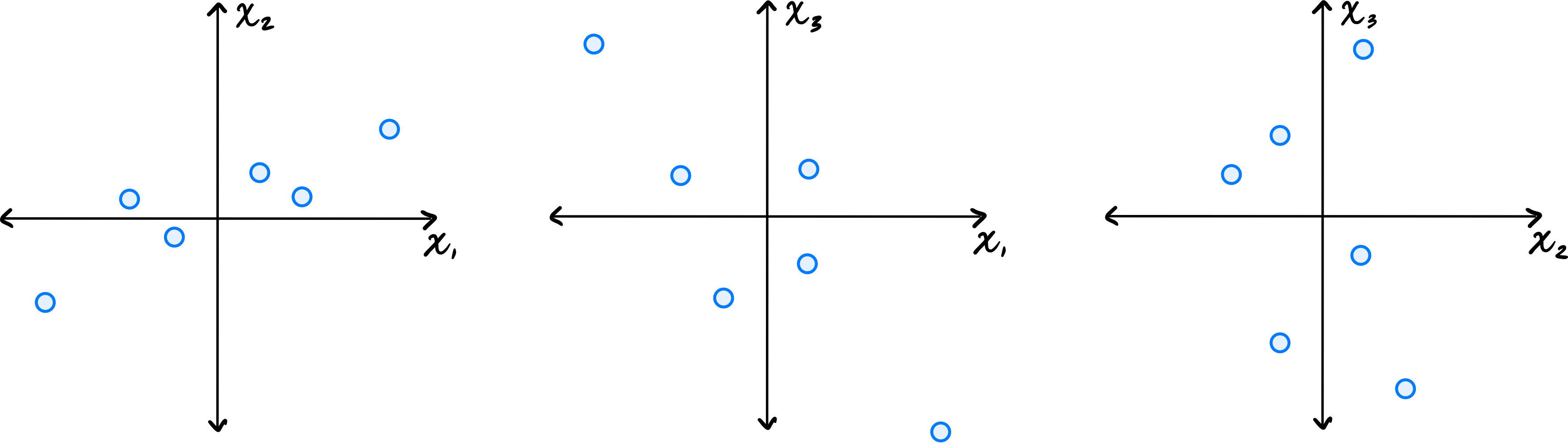
Which one of the following could possibly be the data's sample covariance matrix?
Solution
\(\begin{pmatrix} 10 & 4 & -2 \\ 4 & 5 & 0 \\ -2 & 0 & 10\end{pmatrix}\) From the scatter plots, we can determine the signs of the covariances. The \((x_1, x_2)\) plot shows a positive correlation, so \(C_{12} > 0\). The \((x_1, x_3)\) plot shows a negative correlation, so \(C_{13} < 0\). The \((x_2, x_3)\) plot shows no clear correlation, so \(C_{23}\approx 0\). The only matrix with \(C_{12} > 0\), \(C_{13} < 0\), and \(C_{23} = 0\) is the third choice.
Problem #066
Tags: quiz-04, linear transformations, covariance, lecture-06
Let \(\mathcal X = \{\vec{x}^{(1)}, \ldots, \vec{x}^{(n)}\}\) be a centered data set of \(n\) points in \(\mathbb R^2\). Consider the linear transformation \(\vec f(\vec x) = (-x_1, x_2)^T\), which takes an input point and ``flips'' it over the \(x_2\) axis. Let \(\mathcal X' = \{\vec f(\vec{x}^{(1)}), \ldots, \vec f(\vec{x}^{(n)}) \}\) be the set of points in \(\mathbb R^2\) obtained by applying \(\vec f\) to each point in \(\mathcal X\).
Suppose the sample covariance matrix of \(\mathcal X\) is
What is the sample covariance matrix of \(\mathcal X'\)?
Solution
\(\begin{pmatrix} 5 & 3\\ 3 & 4 \end{pmatrix}\) Flipping the data over the \(x_2\) axis doesn't change the variance (spread) in the \(x_1\) or \(x_2\) directions, and so the diagonal entries of the covariance matrix remain the same. The covariance between \(x_1\) and \(x_2\) stays at the same magnitude but changes sign. That is, the covariance goes from \(-3\) to \(3\).
You can see this more formally, too. The transformation takes a point \((x_1, x_2)^T\) to \((-x_1, x_2)^T\). The covariance between \(x_1\) and \(x_2\) in this transformed data is therefore:
Problem #067
Tags: quiz-04, covariance, variance, lecture-06
Suppose \(C = \begin{pmatrix} 5 & -3\\ -3 & 6 \\ \end{pmatrix}\) is the empirical covariance matrix for a centered data set. What is the variance in the direction given by the unit vector \(\vec{u} = \frac{1}{\sqrt2}(-1, 1)^T\)?
Solution
\(17/2\).
The variance in the direction of a unit vector \(\vec{u}\) is given by \(\vec{u}^T C \vec{u}\).
Problem #068
Tags: covariance, eigenvectors, quiz-04, variance, lecture-06
Let \(C\) be the sample covariance matrix of a centered data set, and suppose \(\vec{u}^{(1)}, \vec{u}^{(2)}, \vec{u}^{(3)}\) are normalized eigenvectors of \(C\) with eigenvalues \(\lambda_1 = 9, \lambda_2 = 3, \lambda_3=0\), respectively.
Suppose \(\vec x = \frac{1}{\sqrt 2}\vec{u}^{(1)} + \frac{1}{\sqrt 6}\vec{u}^{(2)} + \frac{1}{\sqrt 3}\vec{u}^{(3)}\). What is the variance in the direction of \(\vec x\)?
Solution
\(5\) Remember that for any unit vector \(\vec u\), the variance in the direction of \(\vec u\) is given by \(\vec u^T C \vec u\). So, for the vector \(\vec x\), we have
Since \(\vec{u}^{(1)}, \vec{u}^{(2)}, \vec{u}^{(3)}\) are eigenvectors of \(C\), the matrix multiplication simplifies:
Problem #069
Tags: lecture-06, quiz-04, eigenvalues, pca
Let \(C\) be the sample covariance matrix of a centered data set \(\mathcal X\) consisting of five points. Suppose that PCA is performed to reduce the dimensionality of \(\mathcal X\) to one dimension. The results are:
What is the largest eigenvalue of \(C\)?
Solution
\(66/5\) The largest eigenvalue of \(C\) equals the variance of the projected data along the first principal component. This data has mean zero (we can verify: \((4 + 3 - 2 + 1 - 6)/5 = 0\)). So the variance is simply:
Problem #070
Tags: lecture-06, quiz-04, eigenvectors, pca
Consider the data set shown below:
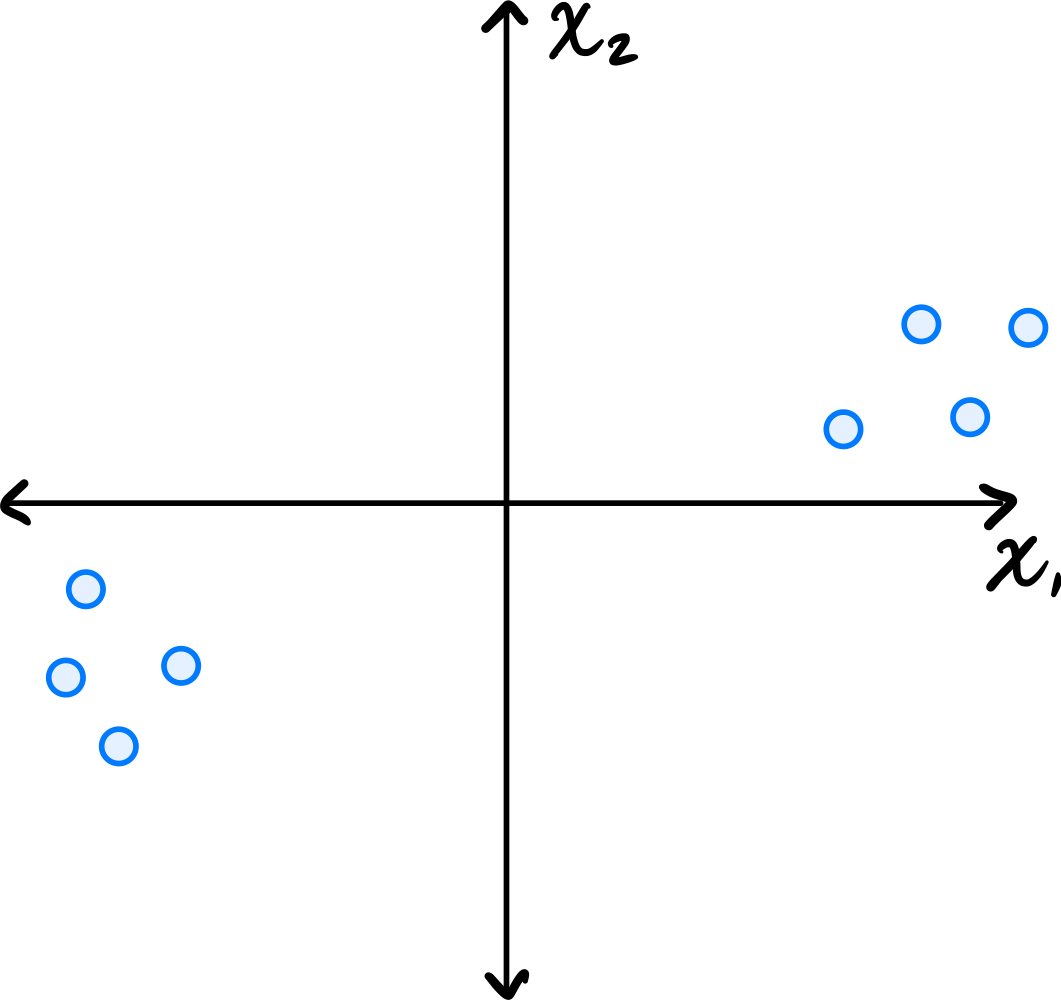
Which of the following could possibly be the top eigenvector of the data's sample covariance matrix?
Solution
\((3, 1)^T\) The top eigenvector of the covariance matrix points in the direction of greatest variance. Looking at the scatter plot, the data is elongated along a direction that has a positive slope, rising more in the \(x_1\) direction than the \(x_2\) direction. The vector \((3, 1)^T\) points roughly in this direction.
\((1, 0)^T\) and \((0, 1)^T\) are along the axes, which don't align with the data's elongation. \((1, 1)^T\) has too steep a slope compared to the apparent direction of the data.
Problem #072
Tags: quiz-04, covariance, variance, lecture-06
Suppose \(C = \begin{pmatrix} 4 & -2\\ -2 & 5 \\ \end{pmatrix}\) is the empirical covariance matrix for a centered data set. What is the variance in the direction given by the unit vector \(\vec{u} = \frac{1}{\sqrt2}(1, 1)^T\)?
Solution
\(5/2\) The variance in the direction of a unit vector \(\vec{u}\) is given by \(\vec{u}^T C \vec{u}\).
Problem #073
Tags: covariance, eigenvectors, quiz-04, variance, lecture-06
Let \(C\) be the sample covariance matrix of a centered data set, and suppose \(\vec{u}^{(1)}, \vec{u}^{(2)}, \vec{u}^{(3)}\) are normalized eigenvectors of \(C\) with eigenvalues \(\lambda_1 = 16, \lambda_2 = 12, \lambda_3=0\), respectively.
Suppose \(\vec x = \frac{1}{\sqrt 2}\vec{u}^{(1)} + \frac{1}{\sqrt 6}\vec{u}^{(2)} + \frac{1}{\sqrt 3}\vec{u}^{(3)}\). What is the variance in the direction of \(\vec x\)?
Solution
\(10\) Remember that for any unit vector \(\vec u\), the variance in the direction of \(\vec u\) is given by \(\vec u^T C \vec u\). So, for the vector \(\vec x\), we have
Since \(\vec{u}^{(1)}, \vec{u}^{(2)}, \vec{u}^{(3)}\) are eigenvectors of \(C\), the matrix multiplication simplifies:
Problem #074
Tags: quiz-04, covariance, lecture-06
Consider the data set in the image shown below:
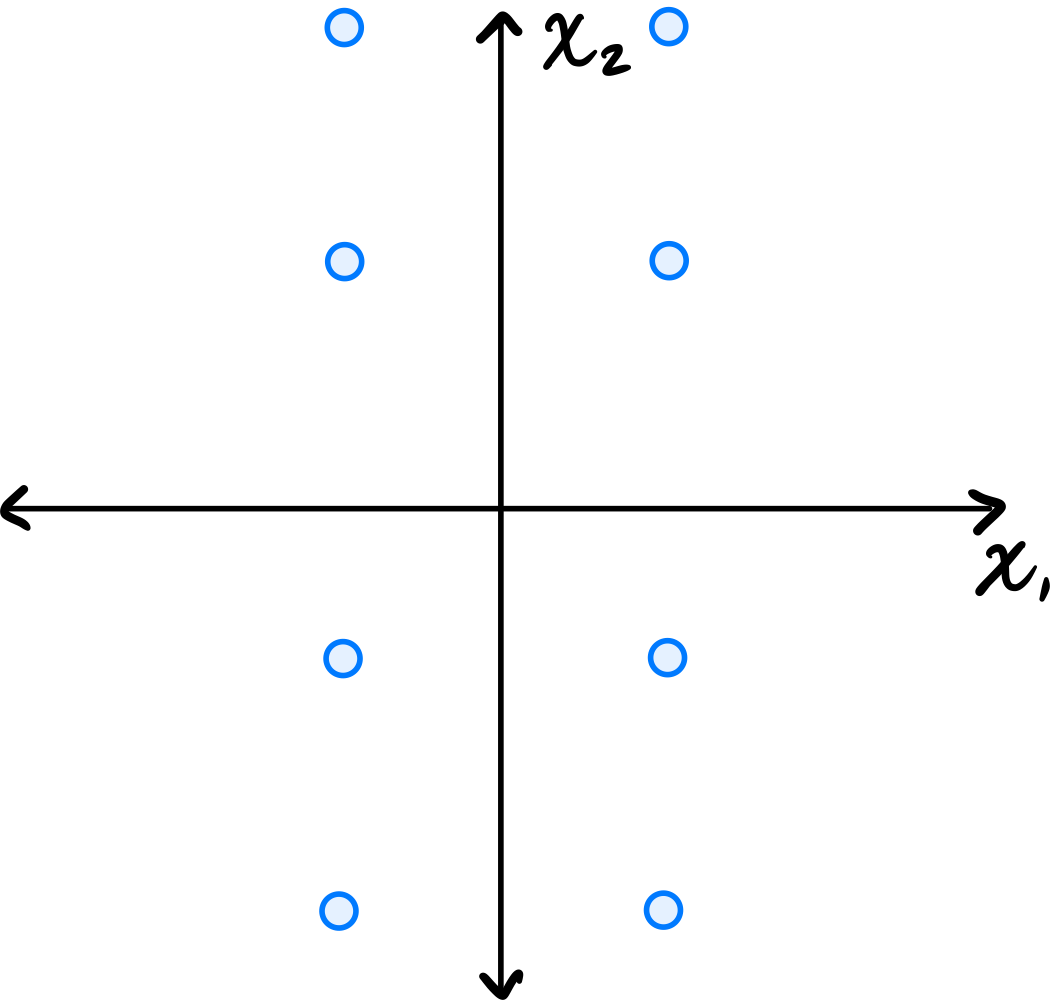
You may assume that this data is already centered and that the symmetry over both the \(x_1\) and \(x_2\) axes is exact.
Consider the \((1, 2)\) entry of the data's sample covariance matrix. Is it positive, negative, or zero?
Solution
It is zero.
Because of the symmetry over both the \(x_1\) and \(x_2\) axes, for every point \((a, b)\) in the data set, there are corresponding points \((-a, b)\), \((a, -b)\), and \((-a, -b)\). When computing the \((1, 2)\) entry of the covariance matrix, we sum \(\tilde{x}_1^{(i)}\cdot\tilde{x}_2^{(i)}\) over all points. For these symmetric groups of four points, the contributions are \(ab\), \(-ab\), \(-ab\), and \(ab\), which sum to zero. Therefore, the \((1, 2)\) entry is zero.
Problem #075
Tags: quiz-04, covariance, lecture-06
Consider the dataset of four points in \(\mathbb R^2\) shown below:
Calculate the sample covariance matrix.
Solution
First, compute the mean:
Then form the centered data matrix \(Z\), whose rows are the centered data points:
The sample covariance matrix is:
Problem #076
Tags: quiz-04, linear transformations, covariance, lecture-06
Let \(\mathcal X = \{\vec{x}^{(1)}, \ldots, \vec{x}^{(n)}\}\) be a data set of \(n\) points in \(\mathbb R^2\). Consider the transformation \(\vec f(\vec x) = (x_1 + 5, x_2 - 5)^T\), which takes an input point and ``shifts'' it over by \(5\) units and down by 5 units. Let \(\mathcal X' = \{\vec f(\vec{x}^{(1)}), \ldots, \vec f(\vec{x}^{(n)}) \}\) be the set of points in \(\mathbb R^2\) obtained by applying \(\vec f\) to each point in \(\mathcal X\).
Suppose the sample covariance matrix of \(\mathcal X\) is
What is the sample covariance matrix of \(\mathcal X'\)?
Solution
\(\begin{pmatrix} 5 & -3\\ -3 & 4 \end{pmatrix}\) The transformation \(\vec f(\vec x) = (x_1 + 5, x_2 - 5)^T\) is a translation (shift) of the data. Translations do not affect the covariance matrix, because in defining the covariance matrix we first subtract the mean from each data point. Translating all points shifts the mean by the same amount, so the centered data remains unchanged.
Problem #077
Tags: lecture-06, quiz-04, eigenvalues, pca
Let \(\mathcal X = \{\vec{x}^{(1)}, \ldots, \vec{x}^{(100)}\}\) be a data set of \(100\) points in 50 dimensions. Let \(\lambda\) be the top eigenvalue of the covariance matrix for \(\mathcal X\).
From this data set, we will construct two data sets \(\mathcal A = \{\vec{a}^{(1)}, \ldots, \vec{a}^{(100)}\}\) and \(\mathcal B = \{\vec{b}^{(1)}, \ldots, \vec{b}^{(100)}\}\) in 25 dimensions, by setting \(\vec{a}^{(i)}\) to be the first 25 coordinates of \(\vec{x}^{(i)}\), and setting \(\vec{b}^{(i)}\) to be the remaining 25 coordinates of \(\vec{x}^{(i)}\).
Let \(\lambda_a\) and \(\lambda_b\) be the top eigenvalues of the covariance matrices for \(\mathcal A\) and \(\mathcal B\), respectively.
True or False: it must be the case that \(\lambda = \max\{\lambda_a, \lambda_b\}\).
Solution
False.
The top eigenvalue \(\lambda\) of the full covariance matrix represents the maximum variance in any direction in the 50-dimensional space. This direction may involve correlations between the first 25 and last 25 coordinates.
When we split the data, \(\lambda_a\) captures only the maximum variance within the first 25 dimensions, and \(\lambda_b\) captures only the maximum variance within the last 25 dimensions. Neither can capture variance in directions that span both coordinate sets.
As a counterexample, consider data where all variance is along the direction \((1, 0, \ldots, 0, 1, 0, \ldots, 0)^T\)(a unit vector with components in both the first 25 and last 25 coordinates). The original data would have \(\lambda > 0\), but both \(\mathcal A\) and \(\mathcal B\) would have smaller top eigenvalues, so \(\lambda > \max\{\lambda_a, \lambda_b\}\).
Problem #078
Tags: pca, dimensionality-reduction, eigenvectors, quiz-04, lecture-06
Let \(C\) be the sample covariance matrix of a data set in \(\mathbb{R}^3\), and suppose \(\vec{u}^{(1)}, \vec{u}^{(2)}, \vec{u}^{(3)}\) are orthonormal eigenvectors of \(C\) with eigenvalues \(\lambda_1 = 9, \lambda_2 = 4, \lambda_3 = 1\), respectively, where:
Suppose a data point is \(\vec{x} = \begin{pmatrix} 3 \\ 6 \\ 6 \end{pmatrix}\).
If PCA is performed to reduce the dimensionality from 3 to 2, what is the new representation of \(\vec{x}\)?
Solution
\(\begin{pmatrix} 8 \\ 4 \end{pmatrix}\) In PCA, to reduce from \(d\) dimensions to \(k\) dimensions, we project each data point onto the top \(k\) eigenvectors (those with the largest eigenvalues).
Here, the top 2 eigenvectors are \(\vec{u}^{(1)}\)(with \(\lambda_1 = 9\)) and \(\vec{u}^{(2)}\)(with \(\lambda_2 = 4\)).
The new representation is obtained by computing the dot product of \(\vec{x}\) with each of the top \(k\) eigenvectors:
Therefore, the new representation is:
Problem #079
Tags: pca, dimensionality-reduction, eigenvectors, quiz-04, lecture-06
Let \(C\) be the sample covariance matrix of a data set in \(\mathbb{R}^4\), and suppose \(\vec{u}^{(1)}, \vec{u}^{(2)}, \vec{u}^{(3)}, \vec{u}^{(4)}\) are orthonormal eigenvectors of \(C\) with eigenvalues \(\lambda_1 = 16, \lambda_2 = 9, \lambda_3 = 4, \lambda_4 = 1\), respectively, where:
Suppose a data point is \(\vec{x} = \begin{pmatrix} 3 \\ 3 \\ 6 \\ 6 \end{pmatrix}\).
If PCA is performed to reduce the dimensionality from 4 to 2, what is the new representation of \(\vec{x}\)?
Solution
\(\begin{pmatrix} 9 \\ 1 \end{pmatrix}\) In PCA, to reduce from \(d\) dimensions to \(k\) dimensions, we project each data point onto the top \(k\) eigenvectors (those with the largest eigenvalues).
Here, the top 2 eigenvectors are \(\vec{u}^{(1)}\)(with \(\lambda_1 = 16\)) and \(\vec{u}^{(2)}\)(with \(\lambda_2 = 9\)).
The new representation is obtained by computing the dot product of \(\vec{x}\) with each of the top \(k\) eigenvectors:
Therefore, the new representation is:
Problem #080
Tags: pca, dimensionality-reduction, eigenvectors, quiz-04, lecture-06
Let \(C\) be the sample covariance matrix of a data set in \(\mathbb{R}^5\), and suppose \(\vec{u}^{(1)}, \vec{u}^{(2)}, \vec{u}^{(3)}, \vec{u}^{(4)}, \vec{u}^{(5)}\) are orthonormal eigenvectors of \(C\) with eigenvalues \(\lambda_1 = 25, \lambda_2 = 16, \lambda_3 = 9, \lambda_4 = 4, \lambda_5 = 1\), respectively, where:
Suppose a data point is \(\vec{x} = \begin{pmatrix} 3 \\ 6 \\ 6 \\ 3 \\ 2 \end{pmatrix}\).
If PCA is performed to reduce the dimensionality from 5 to 3, what is the new representation of \(\vec{x}\)?
Solution
\(\begin{pmatrix} 9 \\ 2 \\ 2 \end{pmatrix}\) In PCA, to reduce from \(d\) dimensions to \(k\) dimensions, we project each data point onto the top \(k\) eigenvectors (those with the largest eigenvalues).
Here, the top 3 eigenvectors are \(\vec{u}^{(1)}\)(with \(\lambda_1 = 25\)), \(\vec{u}^{(2)}\)(with \(\lambda_2 = 16\)), and \(\vec{u}^{(3)}\)(with \(\lambda_3 = 9\)).
The new representation is obtained by computing the dot product of \(\vec{x}\) with each of the top \(k\) eigenvectors:
Therefore, the new representation is:
Problem #081
Tags: lecture-06, dimensionality-reduction, eigenvectors, pca
Consider the following data set of 5 points in \(\mathbb{R}^3\):
Perform PCA on this data set to reduce the dimensionality from 3 to 2. What are the new representations of each data point?
Note: You are not expected to compute the eigenvalues and eigenvectors by hand. Use software (such as numpy.linalg.eigh) to find them.
Solution
First, we form the data matrix and compute the sample covariance matrix:
>>> import numpy as np
>>> X = np.array([
... [3, 1, 2],
... [-1, 2, 0],
... [2, -1, 1],
... [-2, 0, -1],
... [-2, -2, -2]
... ])
>>> mu = X.mean(axis=0)
>>> Z = X - mu
>>> C = 1 / len(X) * Z.T @ Z
Next, we compute the eigendecomposition of \(C\):
>>> eigenvalues, eigenvectors = np.linalg.eigh(C)
>>> idx = eigenvalues.argsort()[::-1]# sort descending
>>> eigenvalues = eigenvalues[idx]
>>> eigenvectors = eigenvectors[:, idx]
>>> eigenvalues
array([6.49, 1.89, 0.01])
To reduce to 2 dimensions, we project onto the top 2 eigenvectors:
>>> U2 = eigenvectors[:, :2]
>>> Z_new = Z @ U2
>>> Z_new
array([[-3.74, -0.13],
[ 0.34, -2.21],
[-1.93, 1.51],
[ 2.16, -0.57],
[ 3.17, 1.40]])
The new representations are (rounded to two decimal places):
Note: The signs of the eigenvectors are not unique; flipping the sign of an eigenvector will flip the sign of the corresponding component in the new representation.
Problem #082
Tags: lecture-06, dimensionality-reduction, eigenvectors, pca
Consider the following data set of 5 points in \(\mathbb{R}^4\):
Perform PCA on this data set to reduce the dimensionality from 4 to 2. What are the new representations of each data point?
Note: You are not expected to compute the eigenvalues and eigenvectors by hand. Use software (such as numpy.linalg.eigh) to find them.
Solution
First, we form the data matrix and compute the sample covariance matrix:
>>> import numpy as np
>>> X = np.array([
... [2, 1, 3, 0],
... [0, -2, 1, 1],
... [-1, 0, -1, 2],
... [1, 2, 0, -1],
... [-2, -1, -3, -2]
... ])
>>> mu = X.mean(axis=0)
>>> Z = X - mu
>>> C = 1 / len(X) * Z.T @ Z
Next, we compute the eigendecomposition of \(C\):
>>> eigenvalues, eigenvectors = np.linalg.eigh(C)
>>> idx = eigenvalues.argsort()[::-1]# sort descending
>>> eigenvalues = eigenvalues[idx]
>>> eigenvectors = eigenvectors[:, idx]
>>> eigenvalues
array([6.35, 2.57, 1.06, 0.02])
To reduce to 2 dimensions, we project onto the top 2 eigenvectors:
>>> U2 = eigenvectors[:, :2]
>>> Z_new = Z @ U2
>>> Z_new
array([[-3.68, 0.43],
[-0.40, -2.19],
[ 0.96, -1.44],
[-0.92, 2.18],
[ 4.03, 1.02]])
The new representations are (rounded to two decimal places):
Note: The signs of the eigenvectors are not unique; flipping the sign of an eigenvector will flip the sign of the corresponding component in the new representation.
Problem #083
Tags: lecture-06, dimensionality-reduction, eigenvectors, pca
Consider the following data set of 5 points in \(\mathbb{R}^4\):
Perform PCA on this data set to reduce the dimensionality from 4 to 3. What are the new representations of each data point?
Note: You are not expected to compute the eigenvalues and eigenvectors by hand. Use software (such as numpy.linalg.eigh) to find them.
Solution
First, we form the data matrix and compute the sample covariance matrix:
>>> import numpy as np
>>> X = np.array([
... [1, 2, 0, 3],
... [-1, 0, 2, 1],
... [2, -1, 1, 0],
... [0, 1, -2, -2],
... [-2, -2, -1, -2]
... ])
>>> mu = X.mean(axis=0)
>>> Z = X - mu
>>> C = 1 / len(X) * Z.T @ Z
Next, we compute the eigendecomposition of \(C\):
>>> eigenvalues, eigenvectors = np.linalg.eigh(C)
>>> idx = eigenvalues.argsort()[::-1]# sort descending
>>> eigenvalues = eigenvalues[idx]
>>> eigenvectors = eigenvectors[:, idx]
>>> eigenvalues
array([5.72, 2.28, 1.34, 0.26])
To reduce to 3 dimensions, we project onto the top 3 eigenvectors:
>>> U3 = eigenvectors[:, :3]
>>> Z_new = Z @ U3
>>> Z_new
array([[-3.45, -1.17, 0.67],
[-1.10, 1.83, 1.02],
[-0.70, 0.83, -2.20],
[ 1.84, -2.31, -0.10],
[ 3.40, 0.82, 0.61]])
The new representations are (rounded to two decimal places):
Note: The signs of the eigenvectors are not unique; flipping the sign of an eigenvector will flip the sign of the corresponding component in the new representation.
Problem #084
Tags: pca, eigenvectors, quiz-04, dimensionality reduction, lecture-06
Suppose \(C\) is a \(3 \times 3\) sample covariance matrix for a data set \(\mathcal X\), and that the top two eigenvectors of \(C\) are:
with eigenvalues \(\lambda_1 = 10\) and \(\lambda_2 = 4\), respectively.
Let \(\vec x = (1, 2, 3)^T\) be the coordinates of \(\vec x\) with respect to the standard basis. Let \(\vec z\) be the result of applying PCA to reduce the dimensionality of \(\vec x\) to 2. What is \(\vec z\)?
Solution
\(\vec z = \left(\frac{1}{\sqrt 2}, \frac{6}{\sqrt 3}\right)^T\).
We compute \(\vec z\) by projecting \(\vec x\) onto each eigenvector:
Problem #086
Tags: pca, eigenvectors, quiz-04, dimensionality reduction, lecture-06
Suppose \(C\) is a \(3 \times 3\) sample covariance matrix for a data set \(\mathcal X\), and that the top two eigenvectors of \(C\) are:
with eigenvalues \(\lambda_1 = 5\) and \(\lambda_2 = 2\), respectively.
Let \(\vec x = (3,2,1)^T\) be the coordinates of \(\vec x\) with respect to the standard basis. Let \(\vec z\) be the result of applying PCA to reduce the dimensionality of \(\vec x\) to 2. What is \(\vec z\)?
Solution
\(\vec z = \left(0, \frac{3}{\sqrt 2}\right)^T\).
We compute \(\vec z\) by projecting \(\vec x\) onto each eigenvector: