Problems tagged with "covariance"
Problem #061
Tags: pca, centering, covariance, quiz-04, lecture-06
Center the data set:
Solution
First, compute the mean:
Then subtract the mean from each data point:
Problem #062
Tags: pca, centering, covariance, quiz-04, lecture-06
Center the data set:
Solution
First, compute the mean:
Then subtract the mean from each data point:
Problem #063
Tags: quiz-04, covariance, lecture-06
Consider the dataset of four points in \(\mathbb R^2\) shown below:
Calculate the sample covariance matrix.
Solution
First, compute the mean:
Then form the centered data matrix \(Z\), whose rows are the centered data points:
The sample covariance matrix is:
Problem #064
Tags: quiz-04, covariance, lecture-06
Consider the data set in the image shown below:
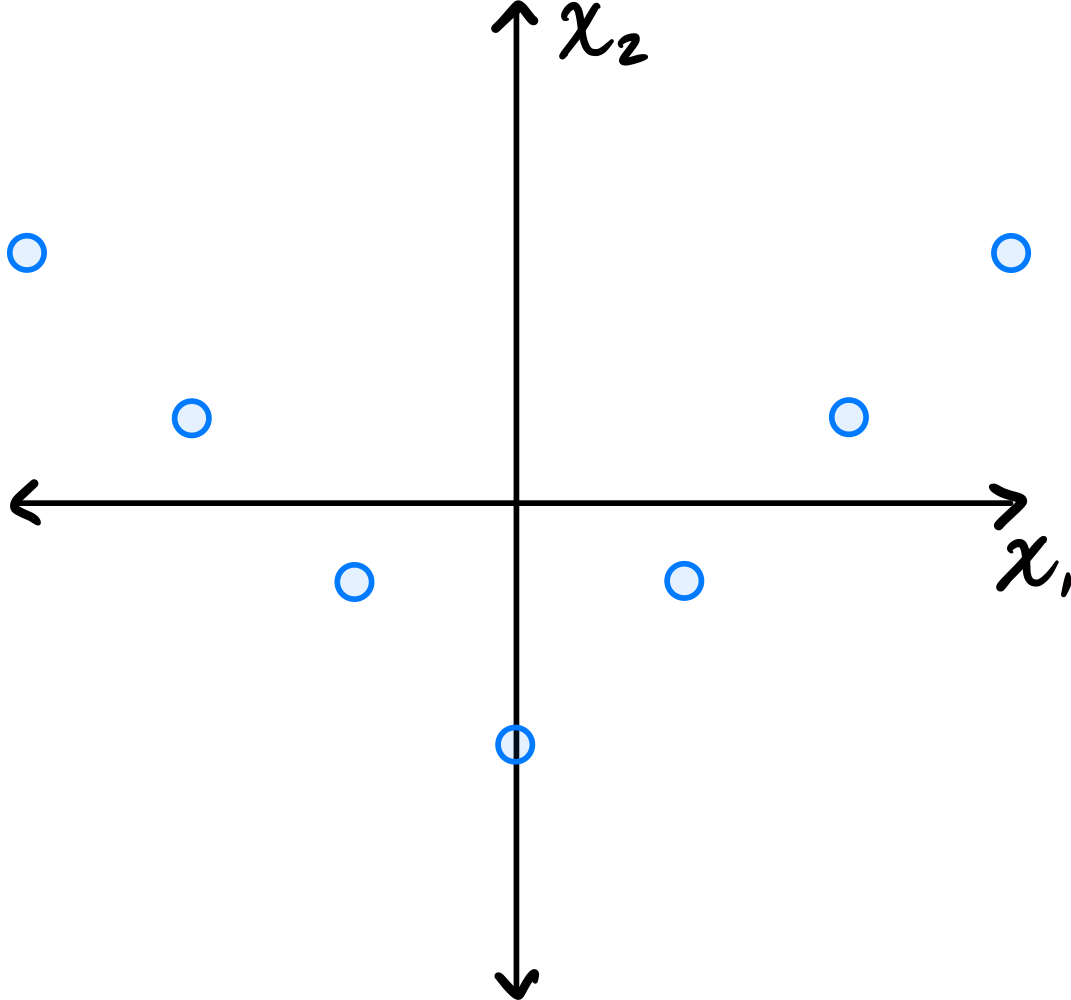
You may assume that this data is already centered and that the symmetry over the \(x_2\) axis is exact.
Which one of the following is true about the \((1, 2)\) entry of the data's sample covariance matrix?
Consider the \((1, 2)\) entry of the covariance matrix, which represents the covariance between the first and second features.
Is the \((1, 2)\) entry of the covariance matrix positive, negative, or zero?
Solution
It is zero.
Because of the symmetry over the \(x_2\) axis, for every point \((a, b)\) in the data set, there is a corresponding point \((-a, b)\). When computing the \((1, 2)\) entry of the covariance matrix, we sum \(\tilde{x}_1^{(i)}\cdot\tilde{x}_2^{(i)}\) over all points. For the symmetric pairs \((a, b)\) and \((-a, b)\), these contributions are \(ab\) and \(-ab\), which cancel out. Therefore, the \((1, 2)\) entry is zero.
Problem #065
Tags: quiz-04, covariance, lecture-06
Let \(\vec{x}^{(1)}, \ldots, \vec{x}^{(6)}\) be a data set of \(6\) points in \(\mathbb R^3\). Shown below are scatter plots of each pair of coordinates (pay close attention to the axis labels):
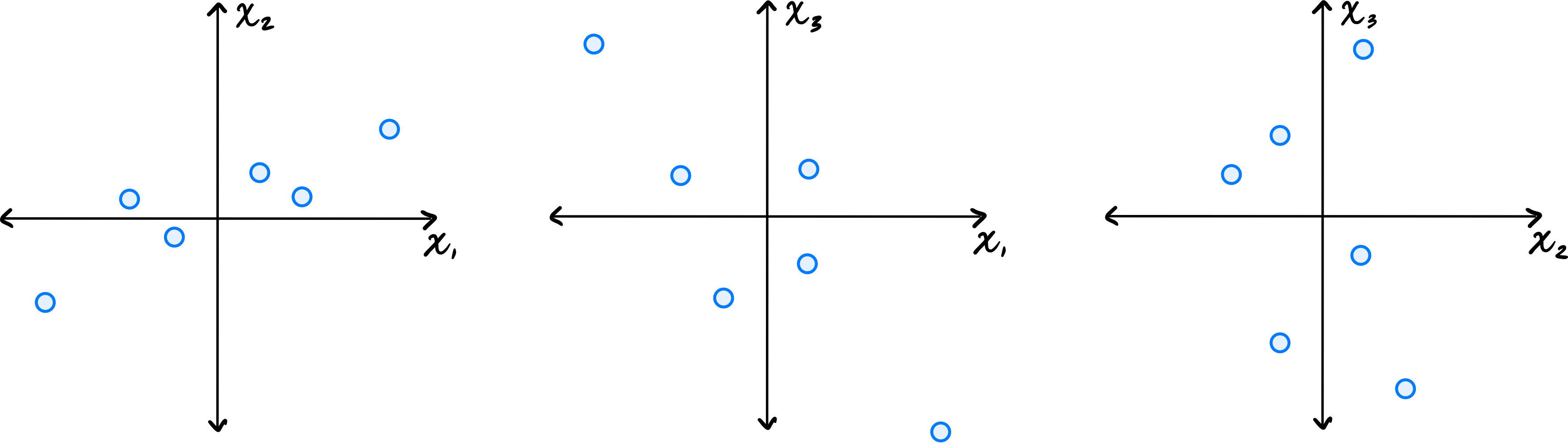
Which one of the following could possibly be the data's sample covariance matrix?
Solution
\(\begin{pmatrix} 10 & 4 & -2 \\ 4 & 5 & 0 \\ -2 & 0 & 10\end{pmatrix}\) From the scatter plots, we can determine the signs of the covariances. The \((x_1, x_2)\) plot shows a positive correlation, so \(C_{12} > 0\). The \((x_1, x_3)\) plot shows a negative correlation, so \(C_{13} < 0\). The \((x_2, x_3)\) plot shows no clear correlation, so \(C_{23}\approx 0\). The only matrix with \(C_{12} > 0\), \(C_{13} < 0\), and \(C_{23} = 0\) is the third choice.
Problem #066
Tags: quiz-04, linear transformations, covariance, lecture-06
Let \(\mathcal X = \{\vec{x}^{(1)}, \ldots, \vec{x}^{(n)}\}\) be a centered data set of \(n\) points in \(\mathbb R^2\). Consider the linear transformation \(\vec f(\vec x) = (-x_1, x_2)^T\), which takes an input point and ``flips'' it over the \(x_2\) axis. Let \(\mathcal X' = \{\vec f(\vec{x}^{(1)}), \ldots, \vec f(\vec{x}^{(n)}) \}\) be the set of points in \(\mathbb R^2\) obtained by applying \(\vec f\) to each point in \(\mathcal X\).
Suppose the sample covariance matrix of \(\mathcal X\) is
What is the sample covariance matrix of \(\mathcal X'\)?
Solution
\(\begin{pmatrix} 5 & 3\\ 3 & 4 \end{pmatrix}\) Flipping the data over the \(x_2\) axis doesn't change the variance (spread) in the \(x_1\) or \(x_2\) directions, and so the diagonal entries of the covariance matrix remain the same. The covariance between \(x_1\) and \(x_2\) stays at the same magnitude but changes sign. That is, the covariance goes from \(-3\) to \(3\).
You can see this more formally, too. The transformation takes a point \((x_1, x_2)^T\) to \((-x_1, x_2)^T\). The covariance between \(x_1\) and \(x_2\) in this transformed data is therefore:
Problem #067
Tags: quiz-04, covariance, variance, lecture-06
Suppose \(C = \begin{pmatrix} 5 & -3\\ -3 & 6 \\ \end{pmatrix}\) is the empirical covariance matrix for a centered data set. What is the variance in the direction given by the unit vector \(\vec{u} = \frac{1}{\sqrt2}(-1, 1)^T\)?
Solution
\(17/2\).
The variance in the direction of a unit vector \(\vec{u}\) is given by \(\vec{u}^T C \vec{u}\).
Problem #068
Tags: covariance, eigenvectors, quiz-04, variance, lecture-06
Let \(C\) be the sample covariance matrix of a centered data set, and suppose \(\vec{u}^{(1)}, \vec{u}^{(2)}, \vec{u}^{(3)}\) are normalized eigenvectors of \(C\) with eigenvalues \(\lambda_1 = 9, \lambda_2 = 3, \lambda_3=0\), respectively.
Suppose \(\vec x = \frac{1}{\sqrt 2}\vec{u}^{(1)} + \frac{1}{\sqrt 6}\vec{u}^{(2)} + \frac{1}{\sqrt 3}\vec{u}^{(3)}\). What is the variance in the direction of \(\vec x\)?
Solution
\(5\) Remember that for any unit vector \(\vec u\), the variance in the direction of \(\vec u\) is given by \(\vec u^T C \vec u\). So, for the vector \(\vec x\), we have
Since \(\vec{u}^{(1)}, \vec{u}^{(2)}, \vec{u}^{(3)}\) are eigenvectors of \(C\), the matrix multiplication simplifies:
Problem #072
Tags: quiz-04, covariance, variance, lecture-06
Suppose \(C = \begin{pmatrix} 4 & -2\\ -2 & 5 \\ \end{pmatrix}\) is the empirical covariance matrix for a centered data set. What is the variance in the direction given by the unit vector \(\vec{u} = \frac{1}{\sqrt2}(1, 1)^T\)?
Solution
\(5/2\) The variance in the direction of a unit vector \(\vec{u}\) is given by \(\vec{u}^T C \vec{u}\).
Problem #073
Tags: covariance, eigenvectors, quiz-04, variance, lecture-06
Let \(C\) be the sample covariance matrix of a centered data set, and suppose \(\vec{u}^{(1)}, \vec{u}^{(2)}, \vec{u}^{(3)}\) are normalized eigenvectors of \(C\) with eigenvalues \(\lambda_1 = 16, \lambda_2 = 12, \lambda_3=0\), respectively.
Suppose \(\vec x = \frac{1}{\sqrt 2}\vec{u}^{(1)} + \frac{1}{\sqrt 6}\vec{u}^{(2)} + \frac{1}{\sqrt 3}\vec{u}^{(3)}\). What is the variance in the direction of \(\vec x\)?
Solution
\(10\) Remember that for any unit vector \(\vec u\), the variance in the direction of \(\vec u\) is given by \(\vec u^T C \vec u\). So, for the vector \(\vec x\), we have
Since \(\vec{u}^{(1)}, \vec{u}^{(2)}, \vec{u}^{(3)}\) are eigenvectors of \(C\), the matrix multiplication simplifies:
Problem #074
Tags: quiz-04, covariance, lecture-06
Consider the data set in the image shown below:
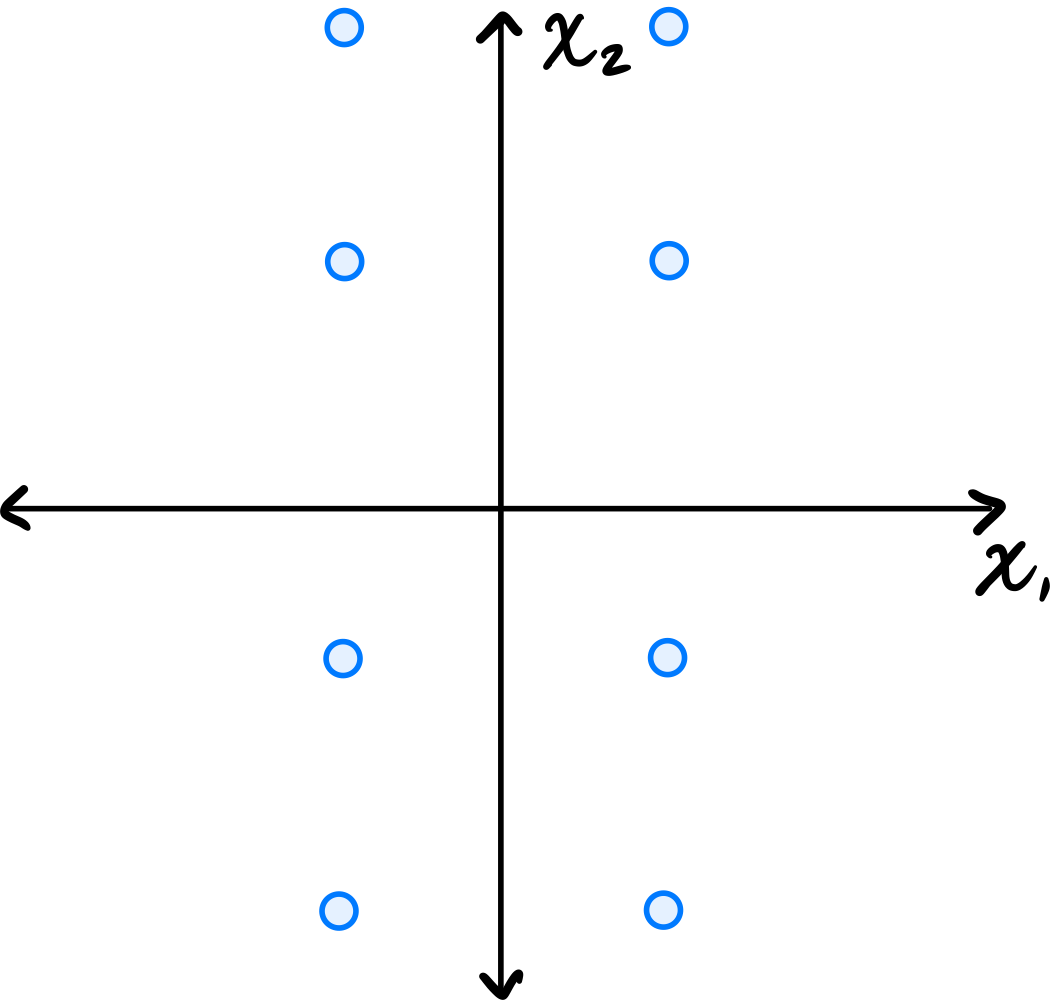
You may assume that this data is already centered and that the symmetry over both the \(x_1\) and \(x_2\) axes is exact.
Consider the \((1, 2)\) entry of the data's sample covariance matrix. Is it positive, negative, or zero?
Solution
It is zero.
Because of the symmetry over both the \(x_1\) and \(x_2\) axes, for every point \((a, b)\) in the data set, there are corresponding points \((-a, b)\), \((a, -b)\), and \((-a, -b)\). When computing the \((1, 2)\) entry of the covariance matrix, we sum \(\tilde{x}_1^{(i)}\cdot\tilde{x}_2^{(i)}\) over all points. For these symmetric groups of four points, the contributions are \(ab\), \(-ab\), \(-ab\), and \(ab\), which sum to zero. Therefore, the \((1, 2)\) entry is zero.
Problem #075
Tags: quiz-04, covariance, lecture-06
Consider the dataset of four points in \(\mathbb R^2\) shown below:
Calculate the sample covariance matrix.
Solution
First, compute the mean:
Then form the centered data matrix \(Z\), whose rows are the centered data points:
The sample covariance matrix is:
Problem #076
Tags: quiz-04, linear transformations, covariance, lecture-06
Let \(\mathcal X = \{\vec{x}^{(1)}, \ldots, \vec{x}^{(n)}\}\) be a data set of \(n\) points in \(\mathbb R^2\). Consider the transformation \(\vec f(\vec x) = (x_1 + 5, x_2 - 5)^T\), which takes an input point and ``shifts'' it over by \(5\) units and down by 5 units. Let \(\mathcal X' = \{\vec f(\vec{x}^{(1)}), \ldots, \vec f(\vec{x}^{(n)}) \}\) be the set of points in \(\mathbb R^2\) obtained by applying \(\vec f\) to each point in \(\mathcal X\).
Suppose the sample covariance matrix of \(\mathcal X\) is
What is the sample covariance matrix of \(\mathcal X'\)?
Solution
\(\begin{pmatrix} 5 & -3\\ -3 & 4 \end{pmatrix}\) The transformation \(\vec f(\vec x) = (x_1 + 5, x_2 - 5)^T\) is a translation (shift) of the data. Translations do not affect the covariance matrix, because in defining the covariance matrix we first subtract the mean from each data point. Translating all points shifts the mean by the same amount, so the centered data remains unchanged.
Problem #085
Tags: reconstruction error, pca, covariance, eigenvalues, lecture-07, quiz-04
Suppose PCA is used to reduce the dimensionality of the centered data shown below from 2 dimensions to 1.
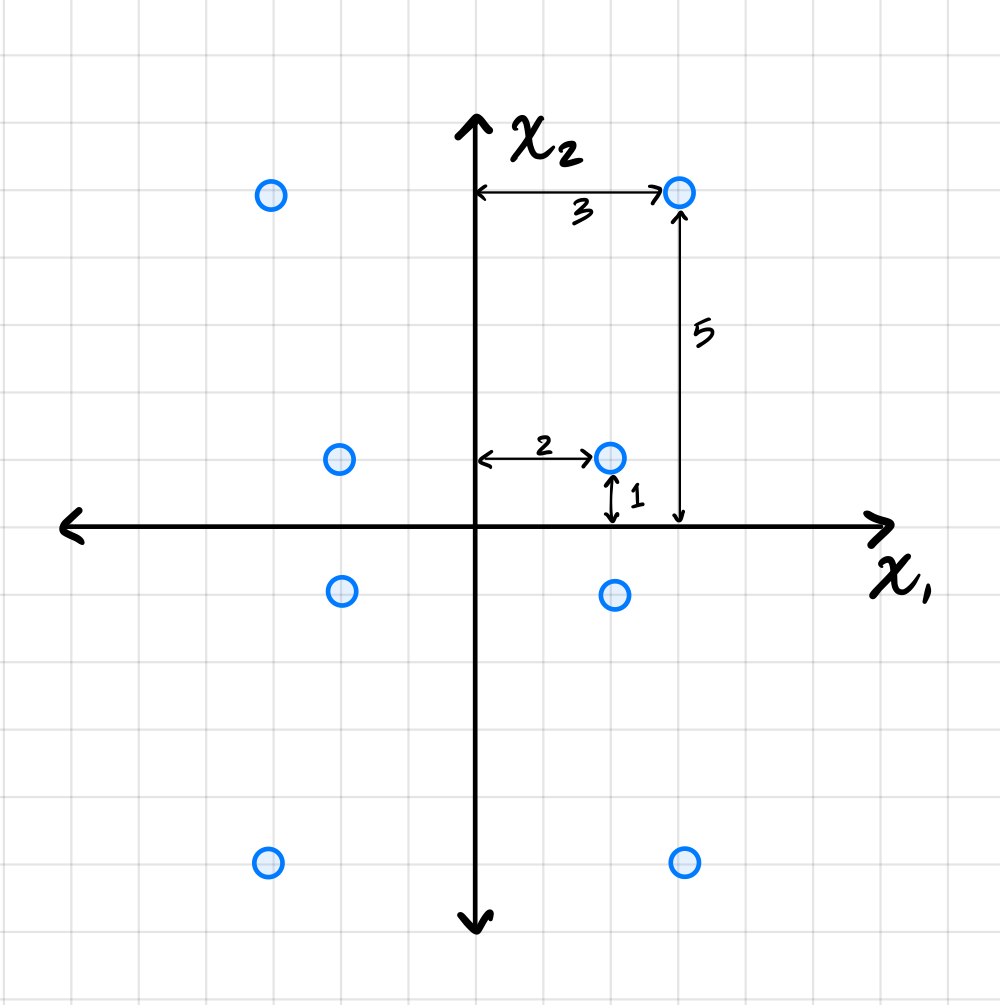
Part 1)
What will be the reconstruction error?
Solution
\(52\).
The first thing to figure out is what the first principal component (first eigenvector of the covariance matrix) is, since this is the direction onto which the data will be projected. Remember that the first eigenvector points in the direction of maximum variance, and in this plot that appears to be straight up (or down). Thus, the first principal component is \((0, 1)^T\)(or \((0, -1)^T\)).
Next, we imagine projecting all of the data onto this line. Since the first eigenvector is vertical, all of the data is projected onto the \(x_2\)-axis. In the process, every point's \(x_1\)-coordinate is lost, and becomes zero, while the \(x_2\)-coordinate remains unchanged. The figure below shows these projected points in red (note that each red point is actually two points on top of each other, since both \((x_1, x_2)^T\) and \((-x_1, x_2)^T\) project to \((0, x_2)^T\)).
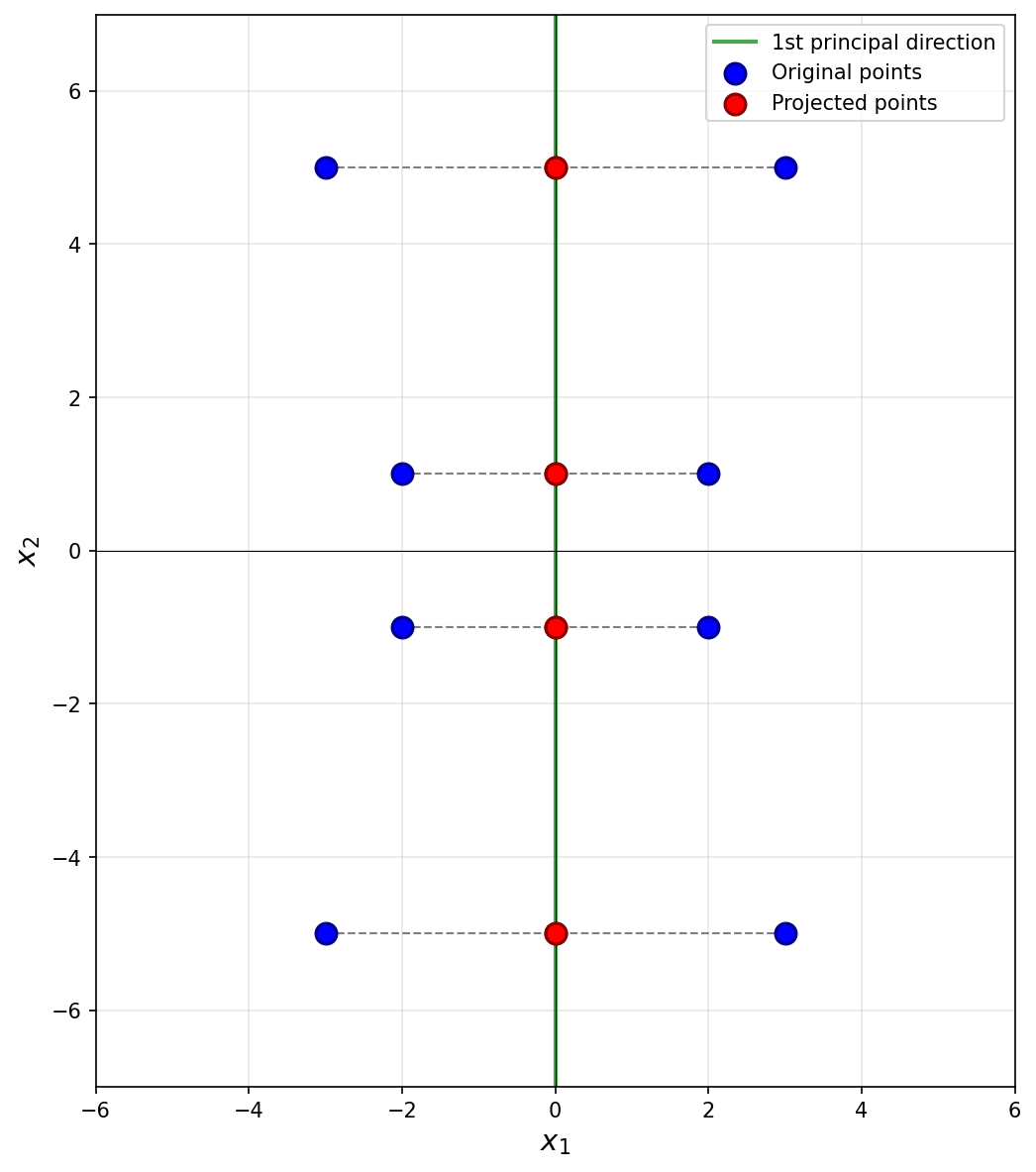
To compute the reconstruction error, we find the squared distance between each point's original position and its projected position. Starting with the upper-right-most point at \((3, 5)^T\), its projection is \((0, 5)^T\), and the squared distance between these two points is \((3 - 0)^2 + (5 - 5)^2 = 9\). For the point at \((2, 1)^T\), its projection is \((0, 1)^T\), and the squared distance is \((2 - 0)^2 + (1 - 1)^2 = 4\).
We could continue, manually calculating the squared distance for each point, but the symmetry of the data allows us to be more efficient. There are four points in the data set that are exactly like the first we just considered and which will have a reconstruction error of 9 each. Similarly, there are four points like the second we considered, each with a reconstruction error of 4. Therefore, the total reconstruction error is:
Part 2)
What is the smallest eigenvalue of the data's covariance matrix?
Solution
\(13/2\).
Remember that the smallest eigenvalue of the data's covariance matrix is equal to the variance of the second PCA feature. That is, it is the variance in the direction of the second principal component (the second eigenvector of the covariance matrix), which in this case is the vector \((1, 0)^T\)(or \((-1, 0)^T\)).
The variance of the data in this direction can be computed by
where \(x_i\) is the \(x_1\)-coordinate of the \(i\)-th data point, \(\mu\) is the mean of all the \(x_1\)-coordinates, and \(n\) is the number of data points. Since hte data is centered, \(\mu = 0\). Therefore, we just need to compute the average of the squared \(x_1\)-coordinates.
Reading these off, we have four points whose \(x_1\)-coordinate is \(3\) or \(-3\), and four points whose \(x_1\)-coordinate is \(2\) or \(-2\). So the variance is:
Therefore, the smallest eigenvalue of the data's covariance matrix is \(13/2\).
Problem #087
Tags: pca, covariance, eigenvalues, lecture-07, quiz-04
Consider the centered data set shown below. The data is symmetric across both axes.
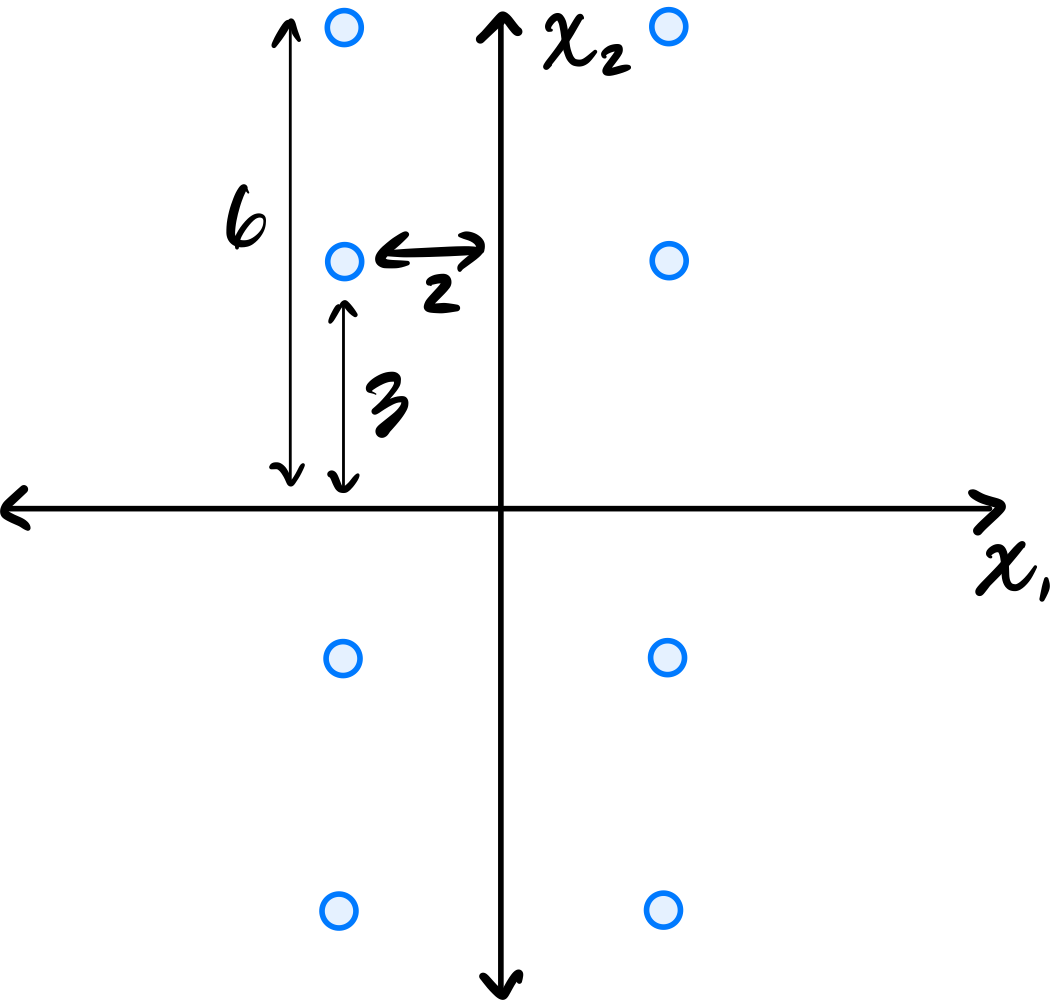
Let \(C\) be the empirical covariance matrix. What is the largest eigenvalue of \(C\)?
Solution
\(45/2\).
Remember that the largest eigenvalue of the data's covariance matrix is equal to the variance of the first PCA feature. That is, it is the variance in the direction of the first principal component (the first eigenvector of the covariance matrix).
Since the data is symmetric across both axes, the eigenvectors of the covariance matrix are aligned with the coordinate axes. From the figure, the data is more spread out vertically than horizontally, so the first principal component is the vector \((0, 1)^T\)(or \((0, -1)^T\)).
The variance of the data in this direction can be computed by
where \(x_i\) is the \(x_2\)-coordinate of the \(i\)-th data point, \(\mu\) is the mean of all the \(x_2\)-coordinates, and \(n\) is the number of data points. Since the data is centered, \(\mu = 0\). Therefore, we just need to compute the average of the squared \(x_2\)-coordinates.
Reading these off, we have four points whose \(x_2\)-coordinate is \(6\) or \(-6\), and four points whose \(x_2\)-coordinate is \(3\) or \(-3\). So the variance is:
Therefore, the largest eigenvalue of the data's covariance matrix is \(45/2\).
Problem #089
Tags: reconstruction error, pca, covariance, eigenvalues, lecture-07, quiz-04
Consider the centered data set consisting of four points:
Suppose PCA is used to reduce the dimensionality of the data from 2 dimensions to 1.
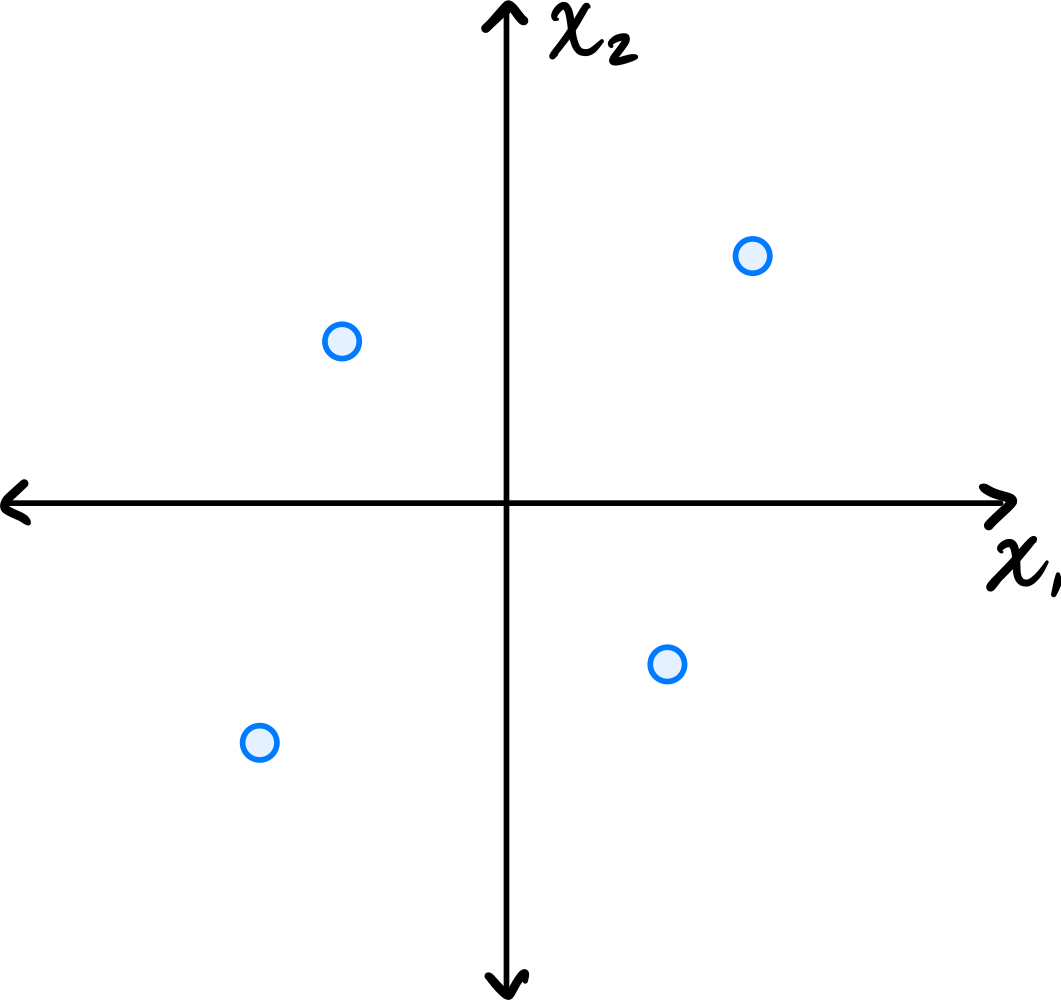
Part 1)
What will be the reconstruction error?
Solution
\(16\).
The first thing to figure out is what the first principal component (first eigenvector of the covariance matrix) is, since this is the direction onto which the data will be projected. Remember that the first eigenvector points in the direction of maximum variance. Looking at the data, the points \((3, 3)^T\) and \((-3, -3)^T\) are farther from the origin than the points \((-2, 2)^T\) and \((2, -2)^T\), so the direction of maximum variance is along the line \(x_2 = x_1\). Thus, the first principal component is \(\frac{1}{\sqrt{2}}(1, 1)^T\)(or \(\frac{1}{\sqrt{2}}(-1, -1)^T\)).
Next, we imagine projecting all of the data onto this line. The figure below shows these projected points in red.
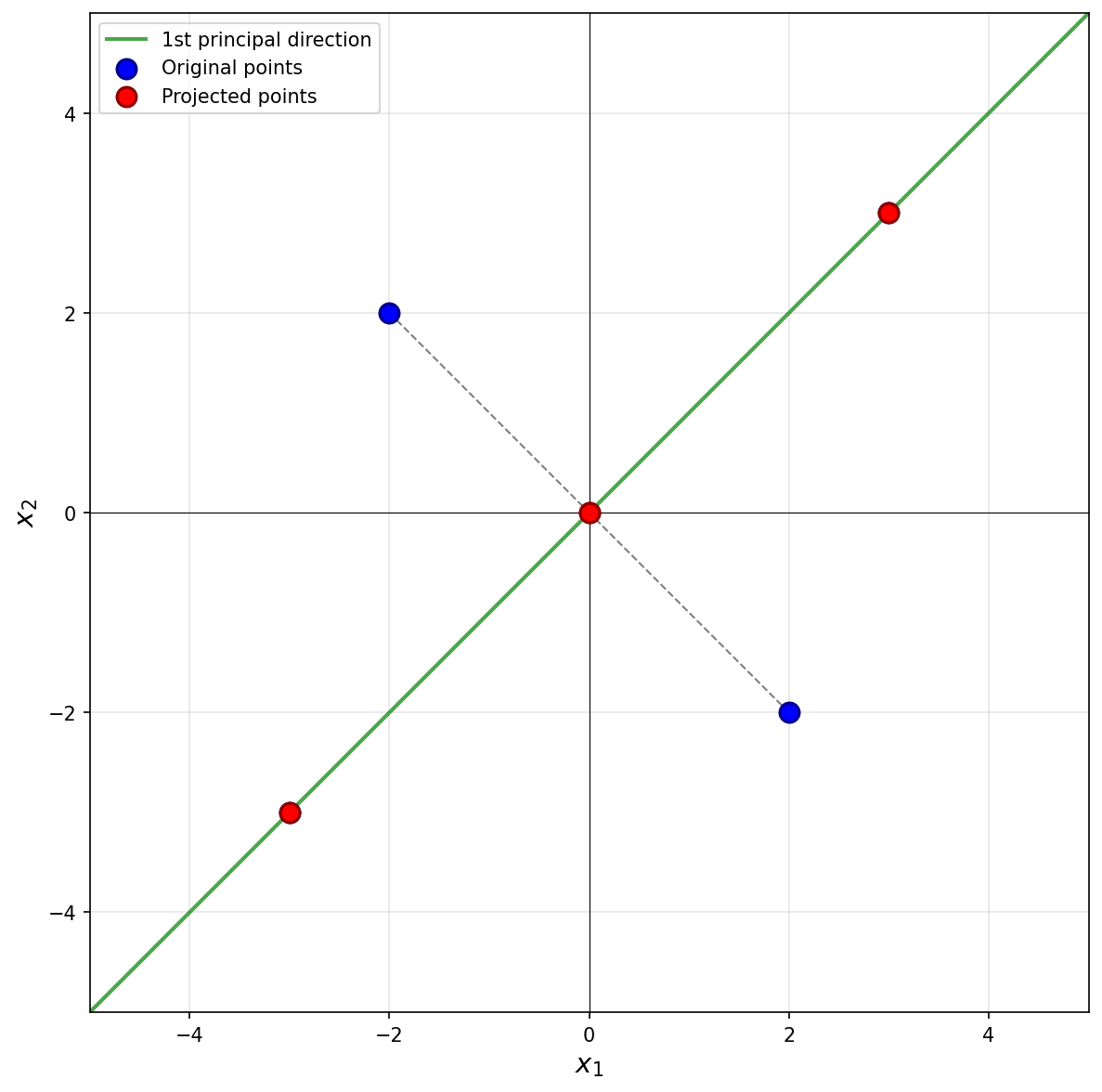
Notice that the points \((3, 3)^T\) and \((-3, -3)^T\) already lie on the line \(x_2 = x_1\), so they project to themselves. Their reconstruction error is zero. The points \((-2, 2)^T\) and \((2, -2)^T\) lie on the line \(x_2 = -x_1\), which is perpendicular to the first principal component. These points both project to the origin \((0, 0)^T\).
To compute the reconstruction error, we find the squared distance between each point's original position and its projected position. For the point at \((-2, 2)^T\), its projection is \((0, 0)^T\), and the squared distance is \((-2 - 0)^2 + (2 - 0)^2 = 4 + 4 = 8\). Similarly, the point \((2, -2)^T\) projects to \((0, 0)^T\) with squared distance \(8\).
Therefore, the total reconstruction error is:
Part 2)
What is the smallest eigenvalue of the data's covariance matrix?
Solution
\(4\).
Remember that the smallest eigenvalue of the data's covariance matrix is equal to the variance of the second PCA feature.
The second PCA feature corresponds to each point's projection onto the second principal component (the second eigenvector of the covariance matrix, and the dashed line in the figure above). In this case, we see that two of the points, \((3, 3)^T\) and \((-3, -3)^T\), project to the origin \((0, 0)^T\), and will have a second PCA feature value of \(0\). The other two points are at \(8\) units away from the origin along this direction, so their second PCA feature values are \(2 \sqrt{2}\) and \(-2 \sqrt{2}\). You could also project these points onto the second principal component to verify this:
Therefore, the second PCA feature values for the four points are 0, 0, 2, and -2. The variance of these (centered) values is: